CRISPR-BASED IMPROVEMENT OF FORAGE DIGESTIBILITY: TARGETING LIGNIN PATHWAYS IN NAPIER GRASS AND MAIZE
DOI:
https://doi.org/10.71146/kjmr903Keywords:
CRISPR/Cas, lignin biosynthesis, forage digestibility, Napier grass, maize, brown midrib, phenylpropanoid pathway, cell-wall recalcitranceAbstract
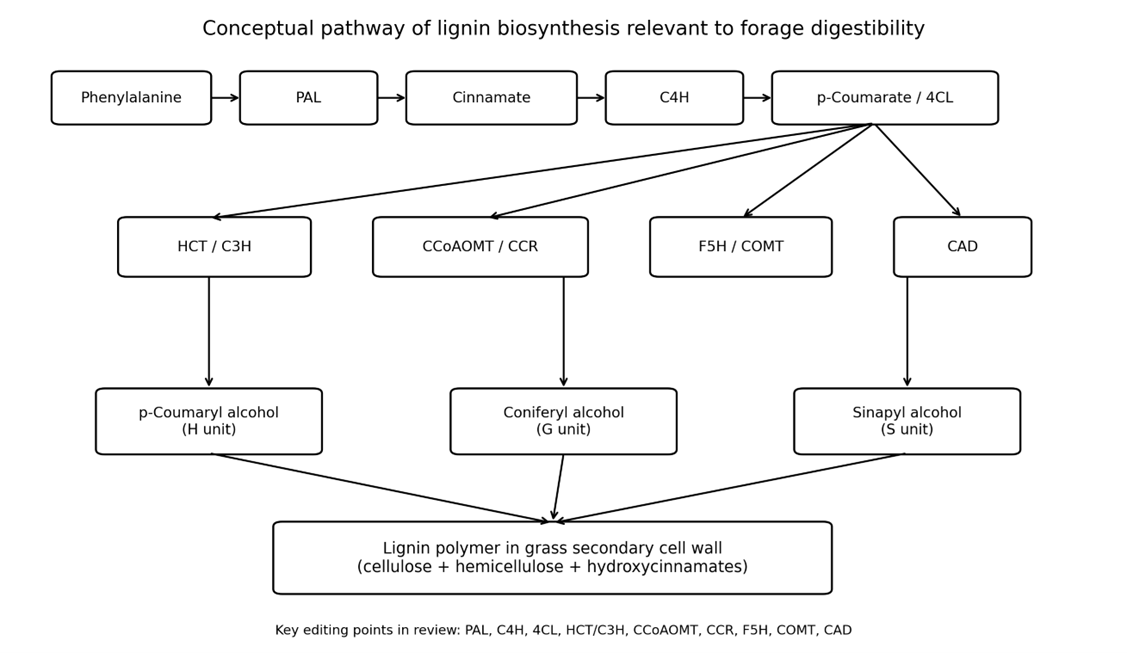
This review examines the potential of CRISPR-based genome editing to improve forage digestibility by modifying lignin biosynthesis in two major C4 grasses, maize (Zea mays L.) and Napier grass (Cenchrus purpureus). Lignin is essential for plant structure, water transport, lodging resistance, and stress adaptation, but it also reduces cell-wall digestibility by limiting microbial and enzymatic access to polysaccharides. In grasses, this challenge is intensified by hydroxycinnamate-mediated cross-linking and the complexity of secondary cell walls. The review synthesizes recent evidence on lignin biosynthetic genes, including PAL, C4H, 4CL, HCT/C3H, CCoAOMT, CCR, F5H, COMT, and CAD, and discusses how editing these targets may alter lignin amount, composition, and cross-linking. A focused comparison is made between maize, where classical brown-midrib mutants and newer genome-editing studies provide a stronger mechanistic foundation, and Napier grass, where genomic resources are improving but functional validation and transformation remain limited. The review methodology is explicitly defined through database searching, screening, and evidence synthesis to address common weaknesses in narrative review writing. Overall, the evidence suggests that moderate, precisely targeted editing of lignin pathways is more promising than drastic lignin reduction, because digestibility gains must be balanced against biomass yield, structural integrity, and stress tolerance. The review concludes that multiplex and regulatory editing strategies, combined with transcriptomic and phenotypic validation, offer the most realistic path for developing high-digestibility forage ideotypes in maize and, in the longer term, Napier grass.
Downloads
References
Liu, Q., Luo, L., & Zheng, L. (2018). Lignin’s: Biosynthesis and biological functions in plants. International Journal of Molecular Sciences, 19(2), 335. https://doi.org/10.3390/ijms19020335
Parachi, L. M., et al. (2024). Grass lignin: biosynthesis, biological roles, and industrial relevance. Frontiers in Plant Science, 15, 1343097. https://doi.org/10.3389/fpls.2024.1343097
Vanhevel, Y., et al. (2024). Breeding for improved digestibility and processing of maize stem cell walls. Frontiers in Plant Science, 15, 1419796. https://doi.org/10.3389/fpls.2024.1419796
Christensen, C. S. L., et al. (2019). Low lignin mutants and reduction of lignin content in grasses for increased utilisation of lignocellulose. Agronomy, 9(5), 256. https://doi.org/10.3390/agronomy9050256
Grabber, J. H. (2004). Genetic and molecular basis of grass cell-wall recalcitrance for forage digestion. Competes Rendos Biologizes, 327(5), 455-465.
Yan, Q., et al. (2021). The elephant grass (Cenchrus purpureus) genome provides insights into anthocyanidin accumulation and fast growth. Plant Biotechnology Journal, 19, 526-541.
Zhang, W., et al. (2020). Tissue-specific transcriptome analysis reveals lignocellulose synthesis regulation in elephant grass (Pennisetum purpureum Schum). BMC Plant Biology, 20, 531. https://doi.org/10.1186/s12870-020-02735-3
Wang, Q., Ruan, M., Li, F., Ma, Z., & Luo, D. (2025). Genome-wide identification and expression analysis of MYB transcription factors involved in lignin biosynthesis in elephant grass (Cenchrus purpureus). Agronomy, 15(6), 1326. https://doi.org/10.3390/agronomy15061326
Negawo, A. T., Teshome, A., Kumar, A., Hanson, J., & Jones, C. S. (2017). Opportunities for Napier grass (Pennisetum purpureum) improvement using molecular genetics. Agronomy, 7(2), 28. https://doi.org/10.3390/agronomy7020028
Namata, M. J., et al. (2025). Genome editing in maize and sorghum. Plant Cell Reports, 44, article information online ahead/issue-dependent pagination.
Wang, Y., et al. (2024). Knockout of ZmNST2 promotes bioethanol production from maize straw by reducing lignin and inhibitory compounds. Biotechnology for Biofuels and Bioproducts, 17, article information online.
Xiong, W., et al. (2020). Characterization of two new brown midrib1 mutations affecting lignin biosynthesis in maize. Plant Physiology, 183, 998-1012.
Wang, S., et al. (2024). Genome-wide identification and characterization of lignin biosynthesis genes in maize. International Journal of Molecular Sciences, 25, 6710. https://doi.org/10.3390/ijms25126710
Yang, Y., et al. (2022). Crop quality improvement through genome editing strategy. Frontiers in Genome Editing, 4, 822810. https://doi.org/10.3389/fgeed.2022.822810
Oliveira, D. M., et al. (2025). CRISPR/Cas9 editing of p-coumaroyl-CoA: monolignol transferase 1 in maize alters phenolic metabolism, lignin structure, and biomass processing. Trends in Plant Science / linked primary report context as indexed online.
Downloads
Published
License
Copyright (c) 2026 Shaiza Sehar, Seerat Fatima, Noreen Sardar, Sayeda Kanza Amina, Sana Bushra, Komal Jaffar (Author)

This work is licensed under a Creative Commons Attribution 4.0 International License.






